| plotMA {limma} | R Documentation |
MA-Plot of Expression Data
Ma Plots Explanation Definition

Description
MA plot and Bland–Altman plot See more » DNA microarray. A DNA microarray (also commonly known as DNA chip or biochip) is a collection of microscopic DNA spots attached to a solid surface. New!!: MA plot and DNA microarray See more » DNA sequencing. DNA sequencing is the process of determining the precise order of nucleotides within a. The MA-plot presents this dye bias even more clearly and also a saturation effect in the Cy5 channel for large intensities. (C) To correct the dye bias, one can perform a local regression (red line) of M (D). The obtained residuals of the local regression, i.e., normalized logged fold changes, are well balanced around zero in MA-plot.
Creates an MA-plot with color coding for control spots.
Usage
Arguments
object | an |
array | integer giving the array to be plotted. |
coef | integer giving the linear model coefficient to be plotted. |
xlab | character string, label for x-axis |
ylab | character string, label for y-axis |
main | character string, title for plot |
status | vector giving the control status of each spot on the array, of same length as the number of rows of |
zero.weights | logical, should spots with zero or negative weights be plotted? |
... | other arguments are passed to |

Details
An MA-plot is a plot of log-intensity ratios (M-values) versus log-intensity averages (A-values).See Ritchie et al (2015) for a brief historical review.
For two color data objects, a within-array MA-plot is produced with the M and A values computed from the two channels for the specified array.This is the same as a mean-difference plot (mdplot) with the red and green log2-intensities of the array providing the two columns.

For single channel data objects, a between-array MA-plot is produced.An artificial array is produced by averaging all the arrays other than the array specified.A mean-difference plot is then producing from the specified array and the artificial array.Note that this procedure reduces to an ordinary mean-difference plot when there are just two arrays total.
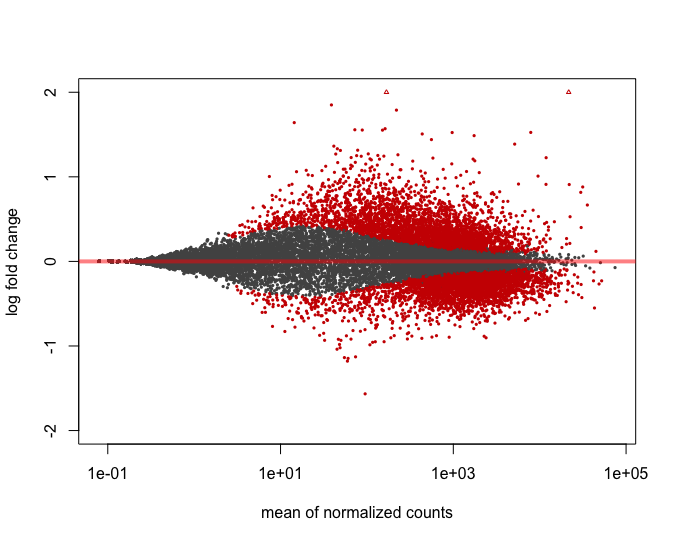
If object is an MArrayLM object, then the plot is an fitted model MA-plot in which the estimated coefficient is on the y-axis and the average A-value is on the x-axis.
The status vector can correspond to any grouping of the probes that is of interest.If object is a fitted model object, then status vector is often used to indicate statistically significance, so that differentially expressed points are highlighted.If object is a microarray data object, then status might distinguish control probes from regular probes so that different types of controls are highlighted.
The status can be included as the component object$genes$Status instead of being passed as an argument to plotMA.
See plotWithHighlights for how to set colors and graphics parameters for the highlighted and non-highlighted points.
Value
A plot is created on the current graphics device.
Note
The plotMD function provides the same functionality as plotMA with slightly different arguments.
Author(s)
Gordon Smyth
References
Ma Plots Explanation
Ritchie, ME, Phipson, B, Wu, D, Hu, Y, Law, CW, Shi, W, and Smyth, GK (2015).limma powers differential expression analyses for RNA-sequencing and microarray studies.Nucleic Acids Research Volume 43, e47.http://nar.oxfordjournals.org/content/43/7/e47
See Also
Ma Plots Explanation Worksheet
The driver function for plotMA is plotWithHighlights.
An overview of plot functions available in LIMMA is given in 09.Diagnostics.